





CENPS rabbit pAb
 One-click to copy product information
One-click to copy product information$148.00/50µL $248.00/100µL
| 50 µL | $148.00 |
| 100 µL | $248.00 |
Overview
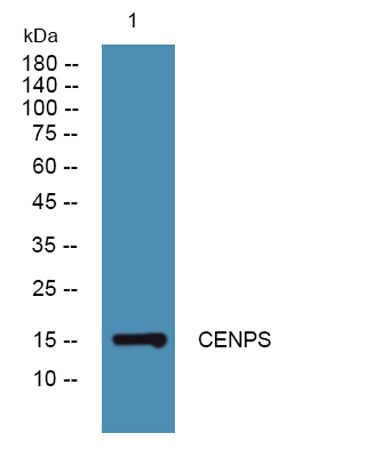
| Product name: | CENPS rabbit pAb |
| Reactivity: | Human;Rat;Mouse; |
| Source: | Rabbit |
| Dilutions: | WB 1:500-2000 ELISA 1:5000-20000 |
| Immunogen: | Synthesized peptide derived from human protein . at AA range: 70-150 |
| Storage: | -20°C/1 year |
| Clonality: | Polyclonal |
| Isotype: | IgG |
| Concentration: | 1 mg/ml |
| Observed Band: | 15kD |
| GeneID: | 100526739 |
| Human Swiss-Prot No: | Q8N2Z9 |
| Cellular localization: | Nucleus . Chromosome, centromere . Chromosome, centromere, kinetochore . Assembly of CENPS and CENPX and its partner subunits CENPT and CENPW at centromeres occurs through a dynamic exchange mechanism. Although exchange is continuous in the cell cycle, de novo assembly starts principally during mid-late S phase and is complete by G2. CENPS is more stably bound at the kinetochore than CENPX (PubMed:19620631, PubMed:24522885). During S phase, rapidly recruited to DNA interstrand cross-links that block replication (PubMed:20347428). Recruited to DNA damage sites about 20 minutes following UV irradiation, reaching a plateau after approximately 40 minutes (PubMed:24522885). . |
| Background: | This gene was identified in the neuroblastoma tumor suppressor candidate region on chromosome 1p36. It contains a TFIID-31 domain, similar to that found in TATA box-binding protein-associated factor, TAF(II)31, which is required for p53-mediated transcription activation. This gene was expressed at very low levels in neuroblastoma tumors, and was shown to reduce cell growth in neuroblastoma cells, suggesting that it may have a role in a cell death pathway. The protein is a component of multiple complexes, including the Fanconi anemia (FA) core complex, the APITD1/CENPS complex, and the CENPA-CAD (nucleosome distal) complex. Known functions include an involvement with chromatin associations of the FA core complex, and a role in the stable assembly of the outer kinetochore. Alternative splicing of this gene results in multiple transcript variants. Naturally occurring read-through transcripts also exist |

 Manual
Manual